Go beyond good enough
Don’t settle for insufficient data hidden behind big brand promises. Go beyond “good enough” to groundbreaking with duet multiomics solutions.
Genetics alone – or conventional 5‑base data – doesn’t tell the full story. A multiomic approach that integrates complete genetics and epigenetics delivers deeper insight into the dynamic state of a cell. DNA methylation at cytosine bases plays a critical role in biological processes, influencing everything from gene expression to cell identity.
Detecting aberrations in methylation patterns, particularly at 5‑methylcytosine (5mC) and its oxidised form 5‑hydroxymethylcytosine (5hmC), is essential. These marks are increasingly recognised as early indicators of disease.
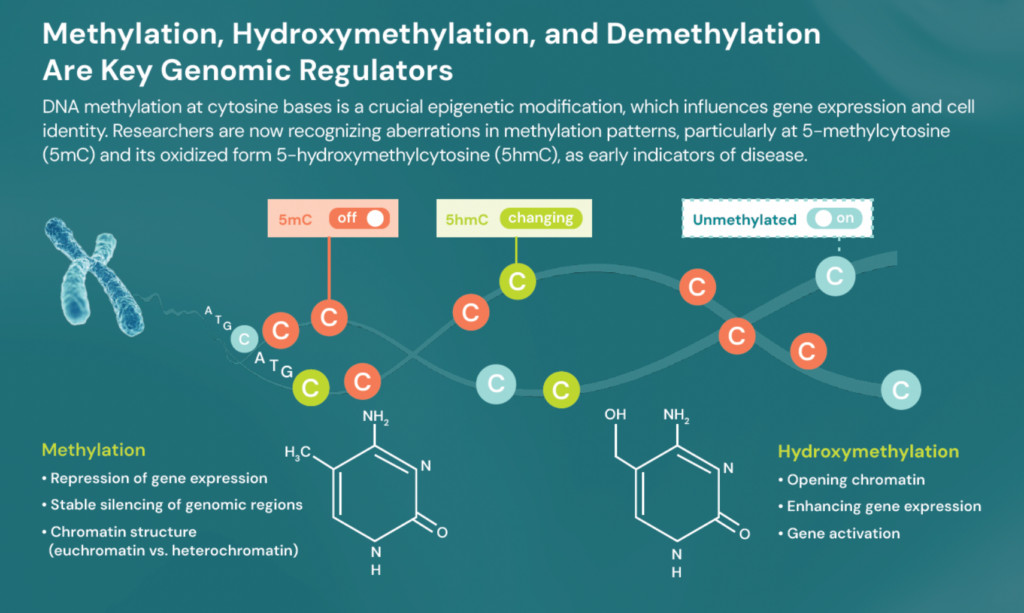
Our advanced 6-base sequencing technology unlocks the epigenetic mechanisms of gene regulation through methylation and hydroxymethylation. Researchers can now accurately measure 5mC and 5hmC, integrated into local genetic context. By capturing multiple layers of biology from a single low-input DNA sample in one experiment, researchers gain comprehensive data to build predictive models of gene expression, chromatin accessibility, and enhancer status – bridging genotype to phenotype.
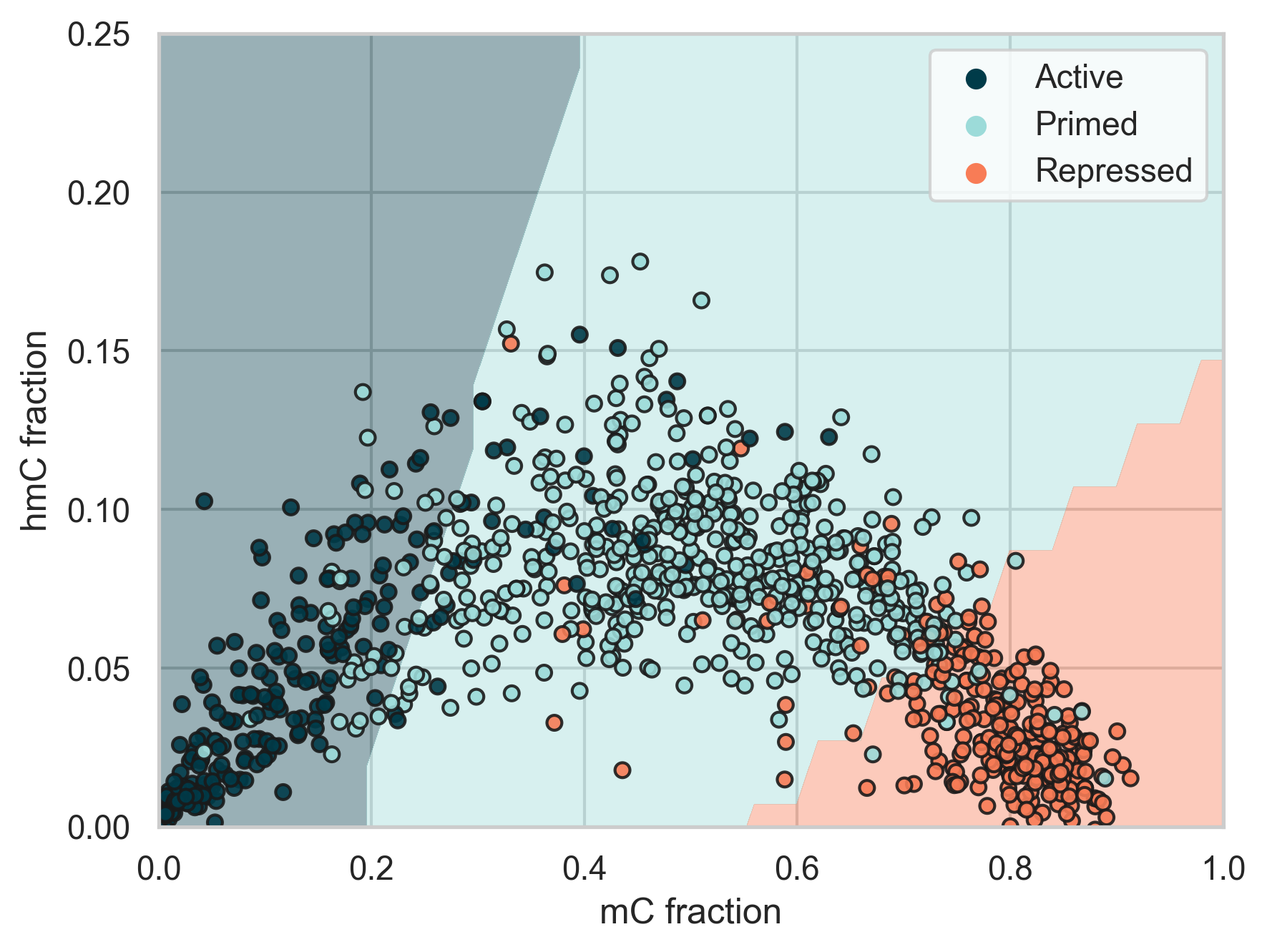
6-base sequencing offers an early window into biological change, revealing genome dynamics at enhancer regions that regulate distal gene expression. Enhancers can be repressed, primed, or active.
Our data show that an increase in 5hmC signals regions committed to activation, transitioning from a high-5mC inactive state to an unmethylated active state.
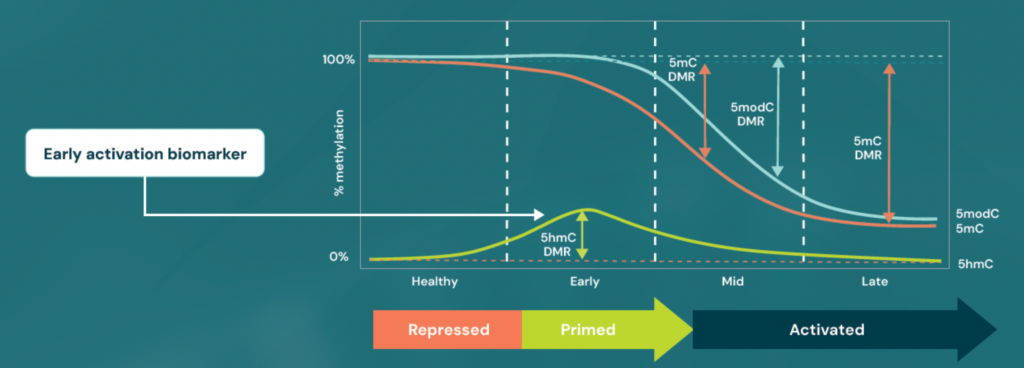
Multiple factors control gene expression, including genetics, epigenetics, and other molecular mechanisms. Specific epigenetic marks can promote (5hmC) or repress (5mC) gene expression. In activating regions, 5mC oxidations to 5hmC primes regions for activation. Conversely, in deactivating regions, 5mC deposition initiates repression. Thus, 5hmC is an early biomarker of activation. When used in conjunction with an increase in 5mC at other regions, the two biomarkers increase the power for early detection.
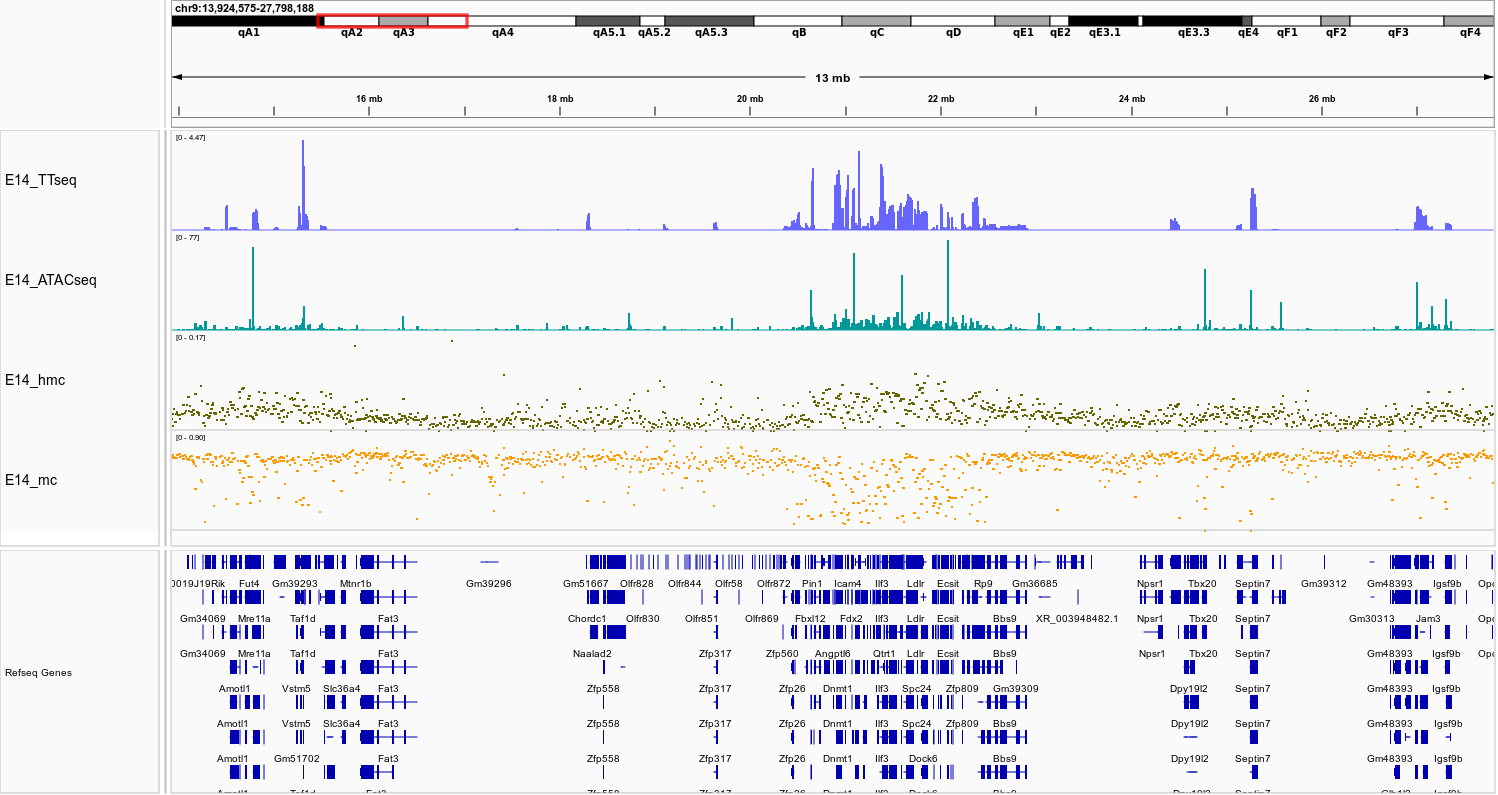
Using the E14 mouse embryonic stem cell model, we plot nascent gene expression (blue), chromatin accessibility (green), 5hmC (brown), and 5mC (orange). Highlighted regions demonstrate how gene expression and chromatin accessibility align with increased 5hmC and reduced 5mC—insights that methylcytosine readouts miss.
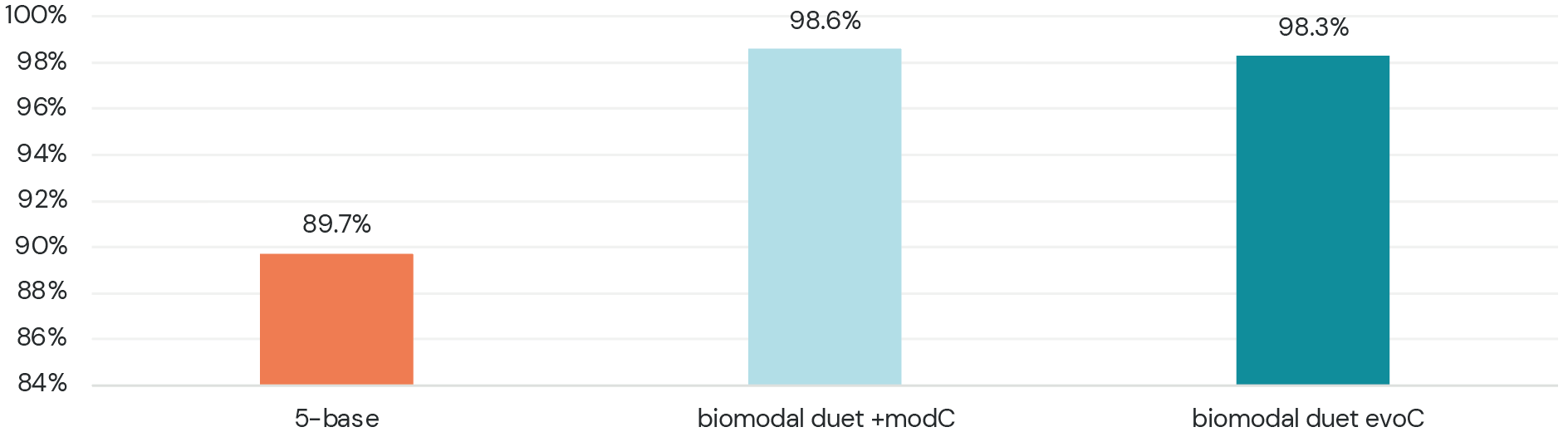
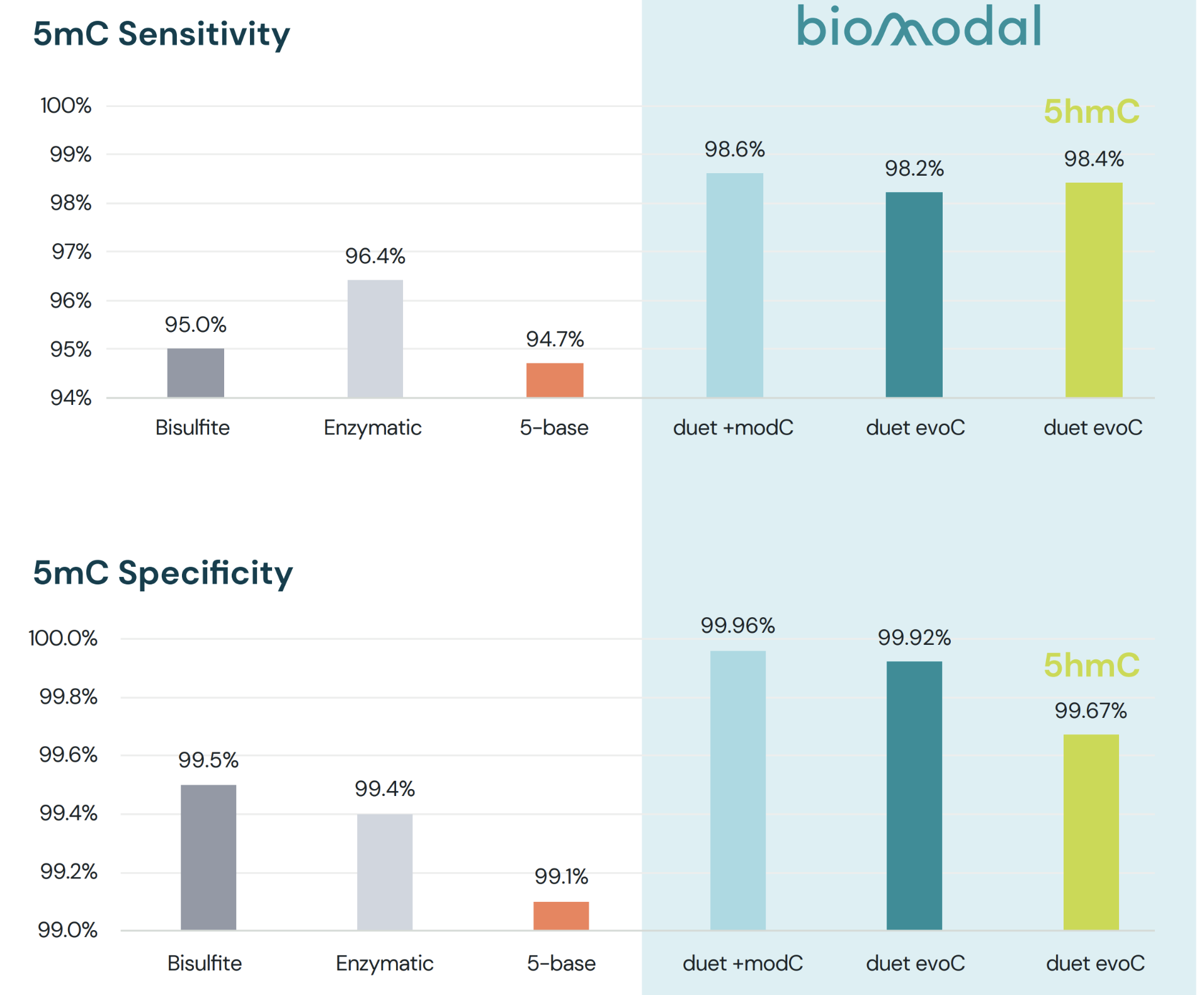
Other 5-base sequencing technologies are missing the mark! Set a new standard.
- A comparison to published data on a 5-base product shows it is less sensitive and less specific than bisulfite
- biomodal’s duet multiomics solution +modC (5-base data) and duet multiomics solution evoC (6-base data) outperform all technologies for 5mC sensitivity and specificity
- biomodal’s duet multiomics solution evoC has the additional advantage of accurately detecting 5hmC as well, an important modification with biological significance
- Modified cytosine is reported when 5mC and 5hmC signals are summed
- biomodal’s duet products separately label and measure 5mC and 5hmC in each run, meaning the sensitivity of the solution to modified cytosine is high
- In comparison, the 5-base product only detects and measures 5mC
- Published data for the 5-base method shows ~5% error rate for calling 5mC
- On average ~5% of modified cytosines in human tissues are 5hmC which are not detected by the 5-base method
- Poor 5mC sensitivity and no 5hmC detection means ~10% of CpGs are missing or miscalled